ʻO ke kaʻina o nā ʻāpana ʻāpana kikoʻī i hoʻonui ʻia (SLAF-Seq)
Nā kikoʻī lawelawe
Papahana ʻenehana
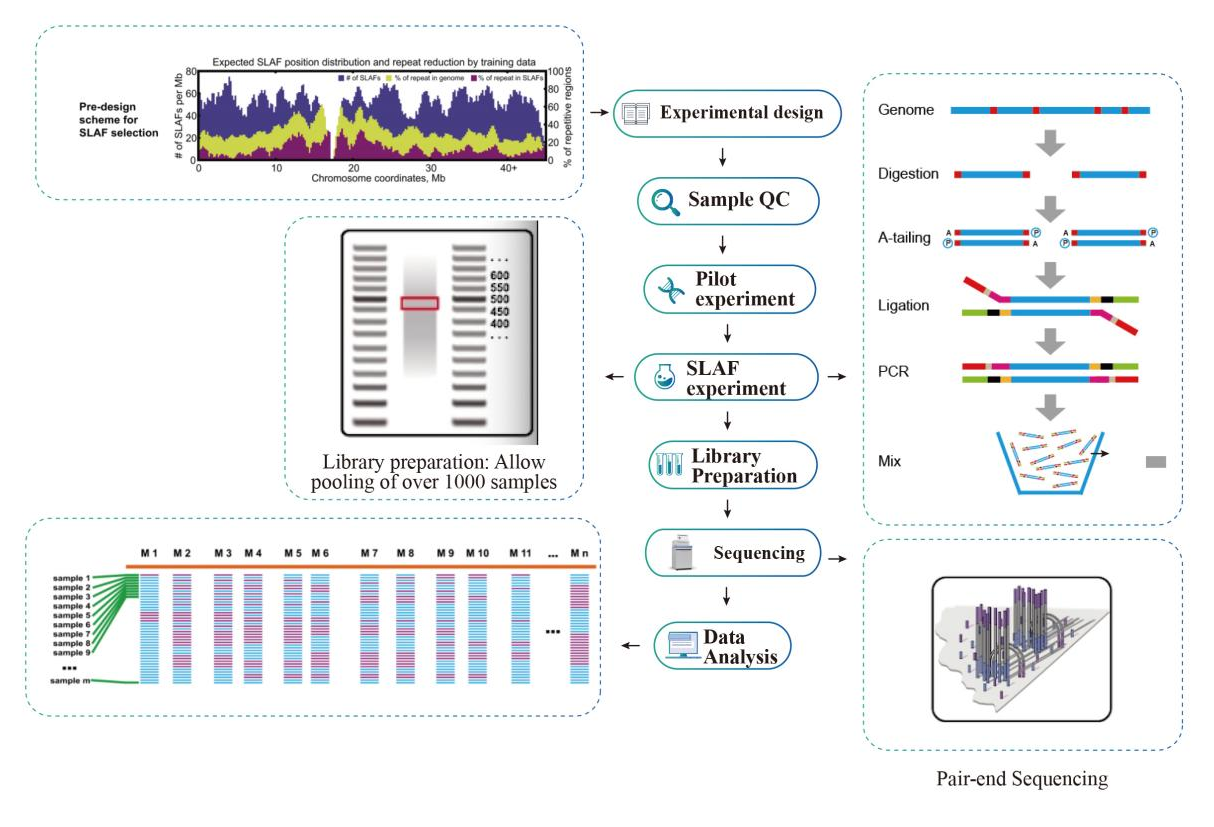
Ka holo hana

Pono lawelawe
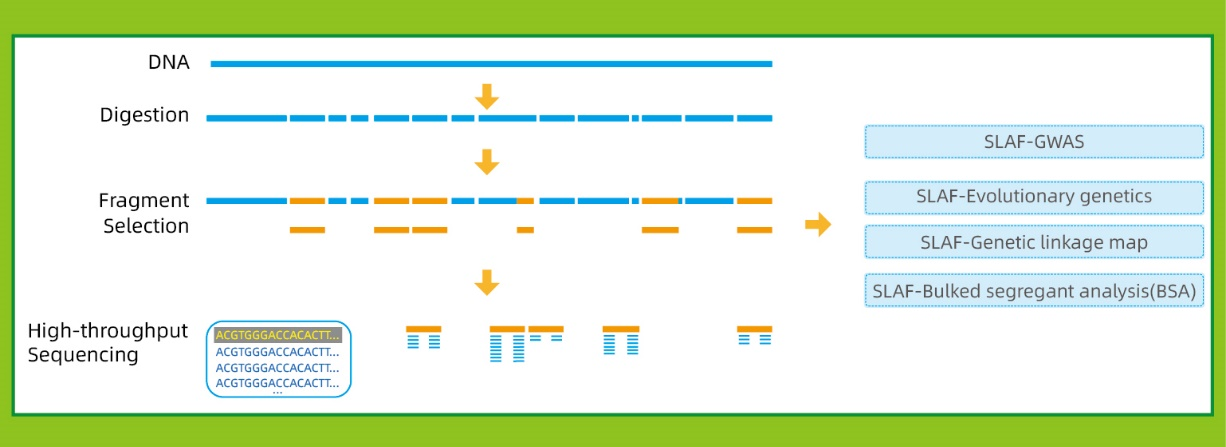
Kiʻekiʻe ka mākaʻikaʻi ʻike ʻana- Ke kōkua nei ka ʻenehana hoʻokaʻina kiʻekiʻe i ka SLAF-Seq i ka ʻike ʻana i nā haneli haneli o nā hōʻailona i loko o ka genome holoʻokoʻa.
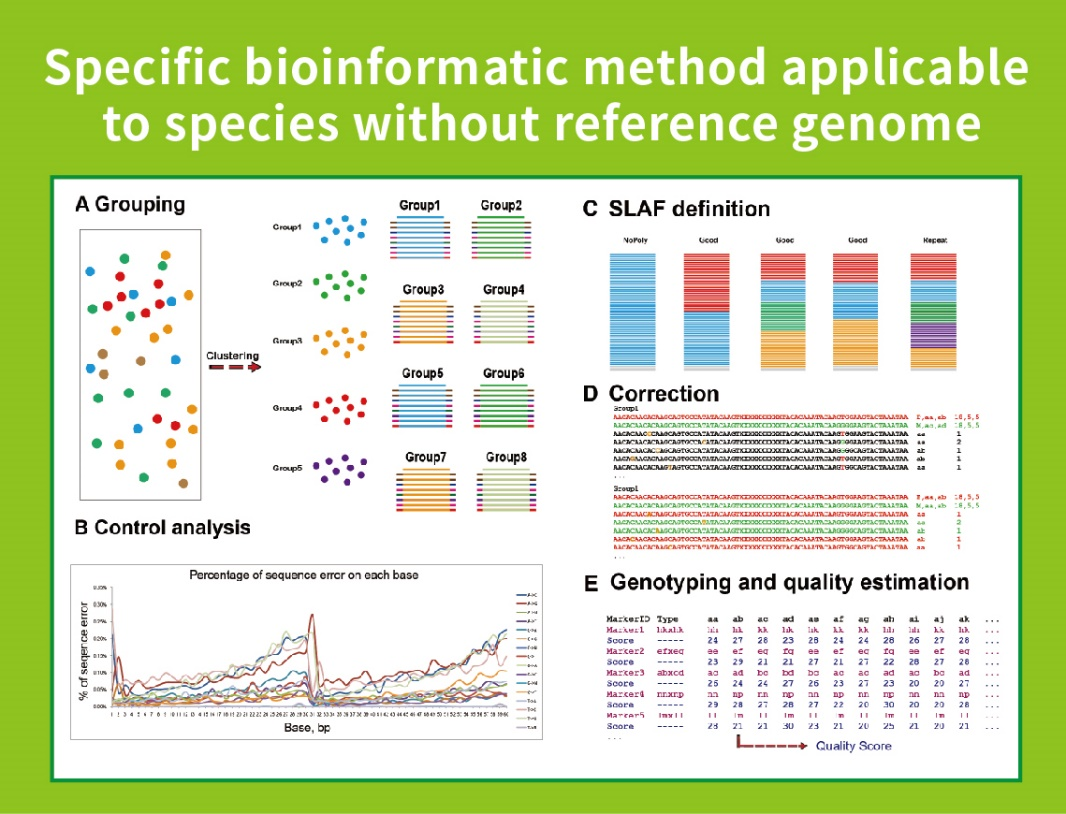
Ka hilinaʻi haʻahaʻa i ka genome- Hiki ke hoʻohana ʻia i nā ʻano me ka ʻole a i ʻole ka genome kuhikuhi.
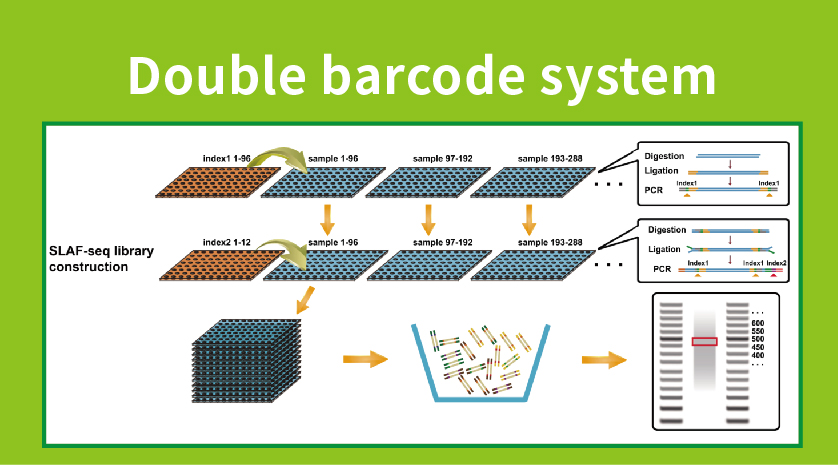
Hoʻolālā papahana maʻalahi- Hoʻokahi-enzyme, dual-enzyme, multi-enzyme digestion a me nā ʻano enzyme like ʻole, hiki ke koho ʻia nā mea āpau e hoʻokō i nā pahuhopu noiʻi ʻokoʻa a i ʻole nā ʻano.Hoʻohana ʻia ka loiloi mua ma ka silico e hōʻoia i kahi hoʻolālā enzyme maikaʻi loa.
ʻO ka digestion enzymatic maikaʻi- Ua hoʻokō ʻia ka hoʻokolohua mua e hoʻopaʻa i nā kūlana, kahi e paʻa ai ka hoʻokolohua maʻamau a hilinaʻi.Hiki ke hoʻokō ʻia ka maikaʻi o ka hōʻiliʻili ʻana ma luna o 95%.
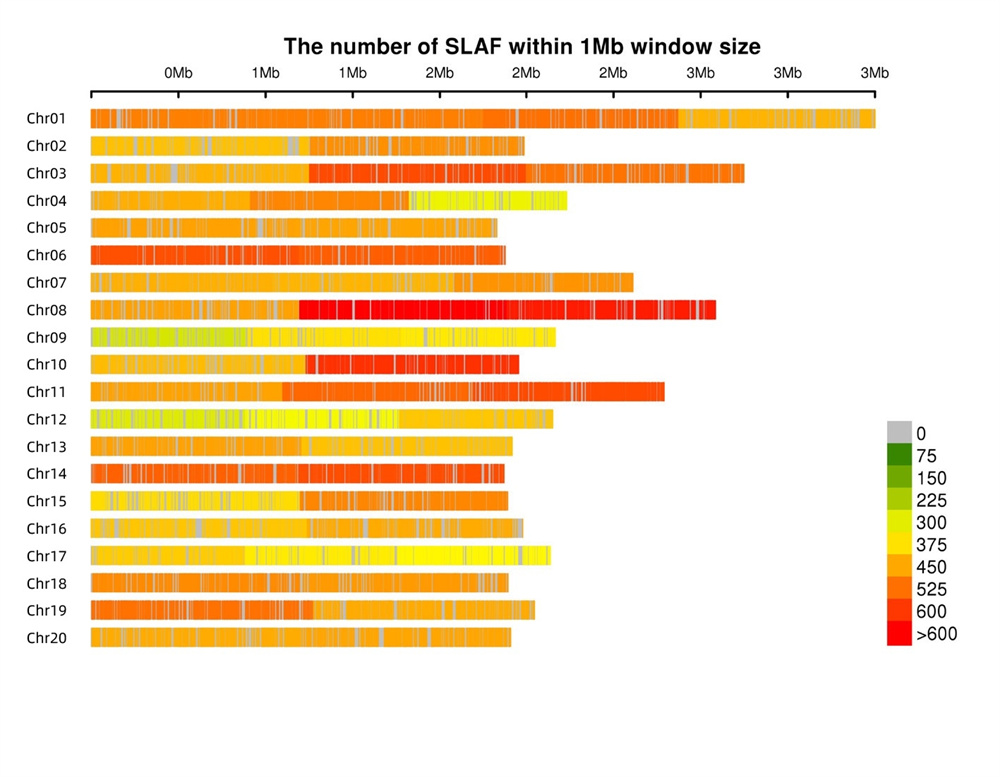
Māhele like ʻia nā hōʻailona SLAF- Hoʻokaʻawale ʻia nā hōʻailona SLAF i nā chromosomes āpau i ka nui loa, e loaʻa ana ka awelika o 1 SLAF no 4 kb.
ʻO ka pale ʻana i ka hana hou ʻana- Hoʻemi ʻia ke kaʻina hana hou i ka ʻikepili SLAF-Seq ma lalo o 5%, ʻoi aku hoʻi i nā ʻano me nā kiʻekiʻe kiʻekiʻe o ka hana hou ʻana, e like me ka palaoa, ka maile, etc.
ʻIke nui- Ma luna o 2000 i pani ʻia nā papahana SLAF-Seq ma nā haneli o nā ʻano e uhi ana i nā mea kanu, nā mammals, nā manu, nā iniseti, nā meaola wai, etc.
ʻO ke kaʻina hana bioinformatic hoʻomohala ponoʻī- Ua hoʻomohala ʻia kahi kaila hana bioinformatic no SLAF-Seq e BMKGENE e hōʻoia i ka hilinaʻi a me ka pololei o ka hopena hope.
Nā kikoʻī lawelawe
| Papahana | Conc.(ng/gl) | Huina (ug) | OD260/280 |
| ʻO Illumina NovaSeq | >35 | >1.6(Volun>15μl) | 1.6-2.5 |
Manaʻo ʻia ʻo Sequencing Strategy
Ka hohonu kaʻina: 10X/Tag
| Nui Genome | Manaʻo ʻia ʻo SLAF Tags |
| < 500 Mb | 100K a i ʻole WGS |
| 500 Mb- 1 Gb | 100 K |
| 1 Gb -2 Gb | 200 K |
| Nā genome nunui a paʻakikī paha | 300 - 400K |
| Nā noi
| Paipai ʻia Ka heluna kanaka
| Hoʻolālā kaʻina a me ka hohonu
| |
| Hohonu
| Helu helu
| ||
| GWAS
| Helu laʻana ≥ 200
| 10X
|
Wahi a ka nui genome
|
| Genetic Evolution
| ʻO kēlā me kēia kanaka pūʻulu liʻiliʻi ≥ 10; huina helu ≥ 30
| 10X
| |
Manaʻo ʻia ka hāʻawi laʻana
ʻO ka pahu: 2 ml centrifuge tube
No ka hapa nui o nā laʻana, paipai mākou ʻaʻole e mālama i loko o ka ethanol.
Laʻana hōʻailona: Pono e hōʻailona ʻia nā laʻana a e like me ka palapala ʻike hāpana i waiho ʻia.
Hoʻouna: Ka hau maloʻo: Pono e hoʻopili mua ʻia nā laʻana i loko o nā ʻeke a kanu ʻia i loko o ka hau maloʻo.
Hana Hana Hana





Laʻana QC
Hoʻokolohua hoʻokele
SLAF-hoʻokolohua
Hoʻomākaukau hale waihona puke
Ke kaʻina ʻana
ʻIkepili ʻikepili
Mahope o ke kuai ana
1. Heluhelu o ka hopena palapala
2. Hoʻomohala ʻana i ka māka SLAF
3. Hōʻike hoʻololi
| Makahiki | Nupepa | IF | Poʻo inoa | Nā noi |
| 2022 | Nā kamaʻilio kūlohelohe | 17.694 | Genomic kumu o ka giga-chromosomes a me ka giga-genome o ka laau peony Paeonia ostii | SLAF-GWAS |
| 2015 | Phytologist hou | 7.433 | Hoʻokumu nā kapuaʻi home i nā wahi genomic koʻikoʻi agronomic i soybeans | SLAF-GWAS |
| 2022 | Nūpepa no ka noiʻi kiʻekiʻe | 12.822 | ʻO ka hoʻokomo ʻana o Gossypium barbadense i loko o G. hirsutum hōʻike i ka loci maikaʻi loa no ka hoʻomaikaʻi like ʻana o ka maikaʻi o ka pulupulu fiber a me ka hua mau ano | SLAF-Evolutionary genetics |
| 2019 | Mea Molekula | 10.81 | Hōʻike ʻia ka heluna kanaka genomic a me ka hui ʻo De Novo i ke kumu o ka weedy ʻO Laiki ma ke ʻano he pāʻani Evolutionary | SLAF-Evolutionary genetics |
| 2019 | Nature Genetics | 31.616 | ʻO ke kaʻina genome a me ka ʻokoʻa genetic o ka carp maʻamau, Cyprinus carpio | Palapala SLAF-Loina |
| 2014 | Nature Genetics | 25.455 | Hāʻawi ka genome o ka pīnī i kanu ʻia i ka ʻike i nā karyotypes legume, polyploid ka hooulu ana a me ka hoohana ana i na mea kanu. | Palapala SLAF-Loina |
| 2022 | Mea Puke Biotechnology Journal | 9.803 | Hōʻike ka ʻike ʻana o ST1 i kahi koho e pili ana i ka hitchhiking o ka morphology hua a me ka aila i loko o ka hana ʻana i ka soya | Hoʻomohala SLAF-Marker |
| 2022 | Nūpepa International o nā ʻepekema Molecular | 6.208 | ʻIke a me ka hoʻomohala ʻana i ka hōʻailona DNA no kahi Wheat-Leymus mollis 2Ns (2D) Hoʻololi Kromoso Disomic | Hoʻomohala SLAF-Marker |