
Osisi/Anụmanụ De Novo Genome Sequencing
Uru ọrụ
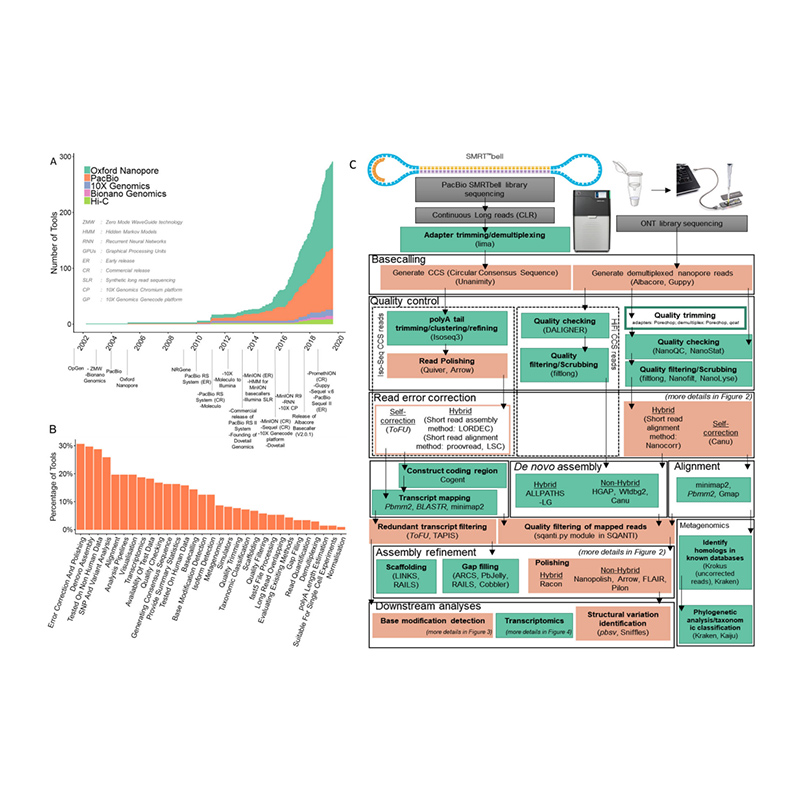
Mmepe nke usoro nhazi usoro na bioinformatics inde novomgbakọ genome
(Amarasinghe SL et al.,Genome Biology, 2020)
● Ịmepụta genome ọhụrụ na imeziwanye mkpụrụ ndụ ihe nketa dị ugbu a maka ụdị mmasị.
● Elu ziri ezi, ịga n'ihu na izu oke na mgbakọ
● Ịmepụta ihe ndị bụ isi maka nyocha n'usoro polymorphism, QTL, ndezi mkpụrụ ndụ ihe nketa, ịzụlite, wdg.
● Ejiri usoro nhazi usoro ọgbọ nke atọ zuru oke: ngwọta mgbakọ genome nkwụsịtụ.
● Usoro na-agbanwe agbanwe na nhazi usoro na-emezu genome dị iche iche nwere atụmatụ dị iche iche
● Ndị otu bioinformatician nwere nkà dị ukwuu nwere ahụmahụ dị ukwuu na mgbakọ genome dị mgbagwoju anya, gụnyere polyploids, genomes giant, wdg.
● Ihe karịrị 100 ikpe na-aga nke ọma nke nwere mmetụta mbipụta nke ihe karịrị narị itoolu
● Ntugharị oge ọsọ ọsọ dịka ọnwa 3 maka mgbakọ genome-ọkwa chromosome.
● Nkwado teknụzụ siri ike nwere usoro patent na ikike nwebisiinka ngwanrọ n'akụkụ nnwale yana bioinformatics.
Nkọwapụta ọrụ
|
Ọdịnaya
|
Platform
|
Gụọ Ogologo
|
Mkpuchi
|
| Nyocha genome
| Illumina NovaSeq
| PE150
| ≥ 50X
|
| Usoro genome
| PacBio Revio
| 15kb HiFi na-agụ
| ≥ 30X
|
| Hi-C
| Illumina NovaSeq
| PE150
| ≥100X
|
Usoro ọrụ

Ihe nlele chọrọ na nnyefe
Ihe atụ chọrọ:
| Ụdị | Anụ ahụ | Maka PacBio | Maka Nanopore |
| Ụmụ anụmanụ | Akụkụ visceral (imeju, splin, wdg) | ≥ 1.0 g | ≥ 3.5 g |
| Akwara | ≥ 1.5 g | ≥ 5.0 g | |
| Ọbara nke mammals | 1.5 ml nke mmiri ara ehi | 5.0 ml nke mmiri ara ehi | |
| Ọbara azụ ma ọ bụ nnụnụ | 0.2 ml nke mmiri ara ehi | 0.5 ml nke mmiri ara ehi | |
| Osisi | Akwụkwọ ọhụrụ | ≥ 1.5 g | ≥ 5.0 g |
| Petal ma ọ bụ azuokokoosisi | ≥ 3.5 g | ≥ 10.0 g | |
| Mgbọrọgwụ ma ọ bụ osisi | ≥ 7.0 g | ≥ 20.0 g | |
| Selụ | Omenala cell | ≥ 3×107 | ≥ 1 × 108 |
Nnyefe nlele akwadoro
Akpa: 2 ml centrifuge tube (Tin foil adịghị akwadoro)
Maka ọtụtụ ihe atụ, anyị na-akwado ka ị ghara ichekwa na ethanol.
Ịkpọ aha nlele: Ọ dị mkpa ka e depụta ihe nlere anya nke ọma yana otu n'ụdị ozi nlele etinyere.
Mbupu: ice-akọrọ: A ga-ebu ụzọ tinye ihe nlele n'ime akpa wee lie ya na ice-akọrọ.
Usoro ọrụ ọrụ

Nnwale imewe
Nbufe ihe atụ
Mwepụta DNA

Ịrụ ụlọ akwụkwọ

Usoro

Nyocha data

Mgbe-ire ọrụ
* Nsonaazụ ngosi egosiri ebe a sitere na genome nke ejiri teknụzụ Biomarker bipụta
1.Circos na chromosome-larịị genome mgbakọ nkeG. rotundifoliumsite Nanopore usoro ikpo okwu
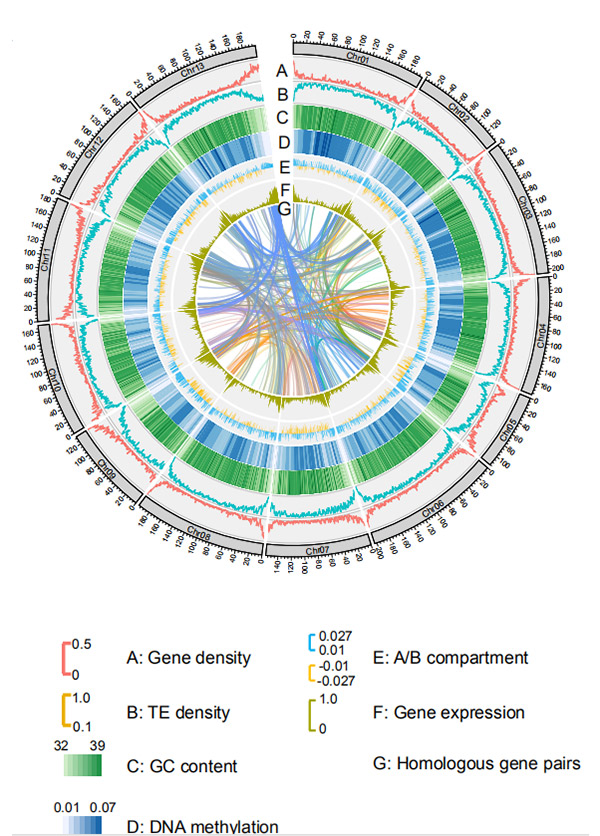
Wang M et al.,Molecular Biology na Evolution, 2021
2.Statistics nke Weining rye genome mgbakọ na nkọwa
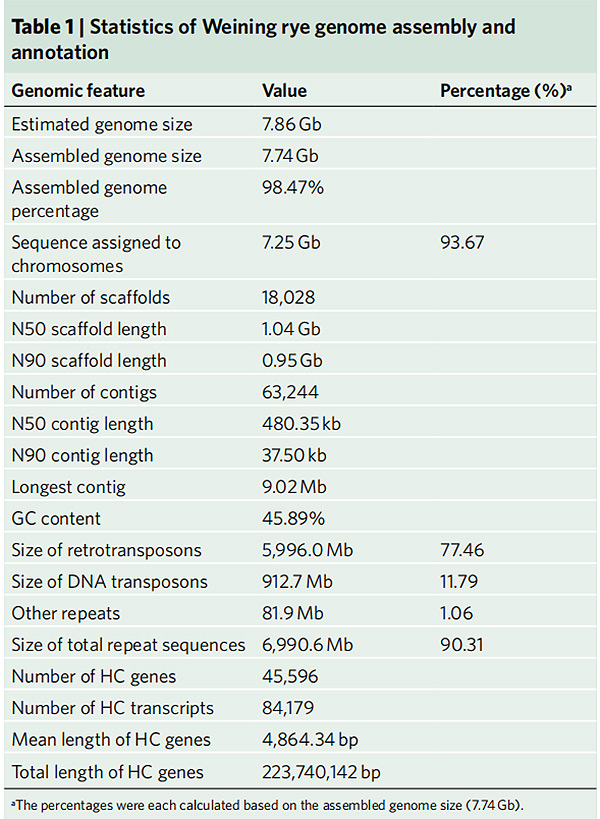
Li G et al.,Genetics okike, 2021
3.Gene amụma nkeSechium edulegenome, sitere na ụzọ amụma atọ:De novoamụma, amụma dabere na Homology na amụma data dabere na RNA-Seq
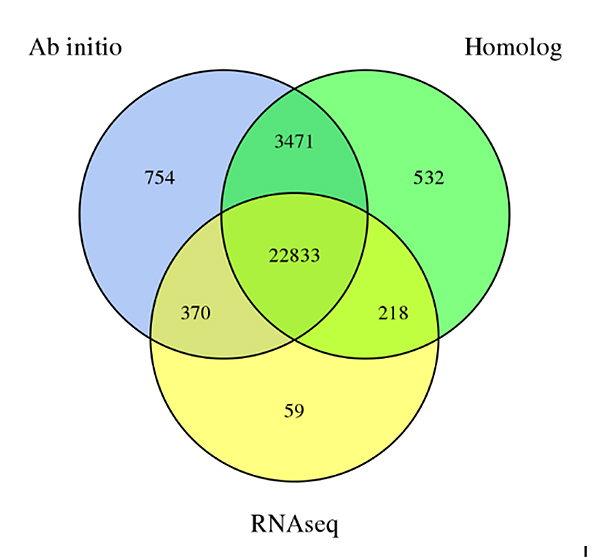
Fu A et al.,Nnyocha Horticulture, 2021
4.Identification nke na-emebibeghị ogologo ọnụ na-ekwughachi na genomes owu atọ
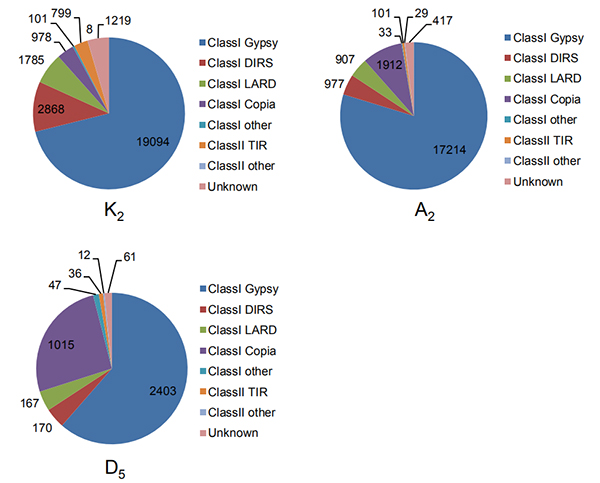
Wang M et al.,Molecular Biology na Evolution, 2021
5.Hi-C okpomọkụ map nkeC. acuminatagenome na-egosi mmekọrịta genome-obosara niile.Ike nke mmekọrịta Hi-C dabara na anya ahịrị n'etiti contigs.Ahịrị kwụ ọtọ dị ọcha na maapụ okpomoku a na-egosi njigide contigs nke ọma na chromosomes.(Nha na-aga n'ihu: 96.03%)
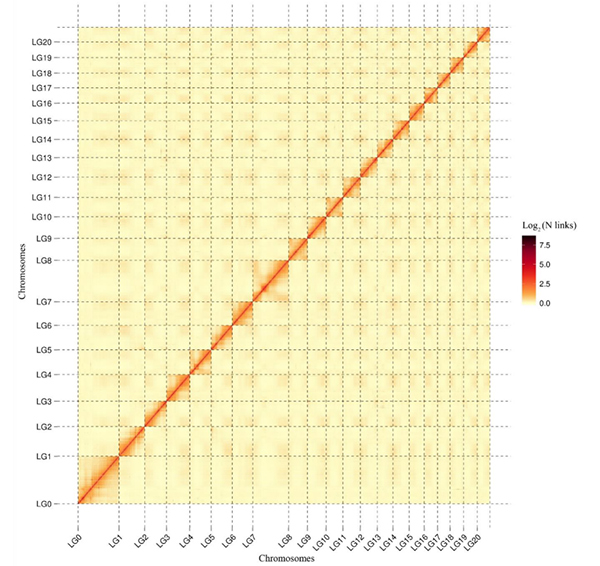
kang M et al.,Nkwukọrịta okike,2021
Okwu BMK
Mgbakọ genome dị elu na-egosipụta njirimara genomic rye na mkpụrụ ndụ ihe nketa dị mkpa
Ebipụtara: Genetics okike, 2021
Atụmatụ usoro:
Mgbakọ Genome: Ụdị PacBio CLR nwere ọbá akwụkwọ 20 kb (497 Gb, ihe dị ka 63 ×)
Ndozi usoro: NGS nwere ọbá akwụkwọ DNA 270 bp (430 Gb, ihe dị ka 54 ×) n'elu ikpo okwu Illumina
Nkwụchi aka: Ọbá akwụkwọ Hi-C (560 Gb, ihe dị ka 71 ×) n'elu ikpo okwu Illumina
Maapụ anya: (779.55 Gb, ihe dị ka 99 ×) na Bionano Irys
Nsonaazụ igodo
1.E bipụtara mgbakọ nke Weining rye genome na ngụkọta genome nke 7.74 Gb (98.74% nke atụmatụ genome nha site na cytometry eruba).Scaffold N50 nke mgbakọ a nwetara 1.04 Gb.93.67% nke contigs enwetara nke ọma na 7 pseudo-chromosomes.A tụlere ọgbakọ a site na map njikọ, LAI na BUSCO, nke butere nnukwu akara na nyocha niile.
2.Ọmụmụ ihe ndị ọzọ gbasara genomics comparative, genetic linkage map, transcriptomics ọmụmụ ka emere na ndabere nke genome a.Ekpughere usoro ihe omume metụtara mkpụrụ ndụ ihe nketa gụnyere genome-wide gene mbiputegharị na mmetụta ha na mkpụrụ ndụ ihe nketa starch biosynthesis;nhazi anụ ahụ nke prolamin loci gbagwojuru anya, njirimara mkpụrụ ndụ ihe nketa na-adabere n'ụdị isi mmalite yana mpaghara chromosomal nke jikọtara ụlọ na loci na rye.
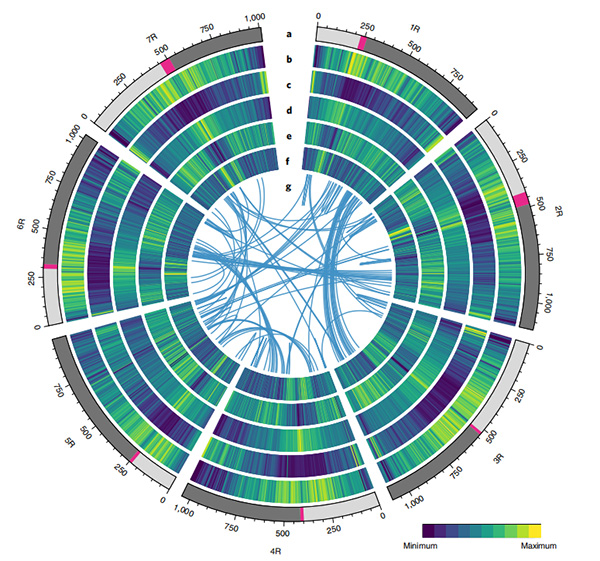 Ihe osise Circos na njirimara genomic nke Weining rye genome | 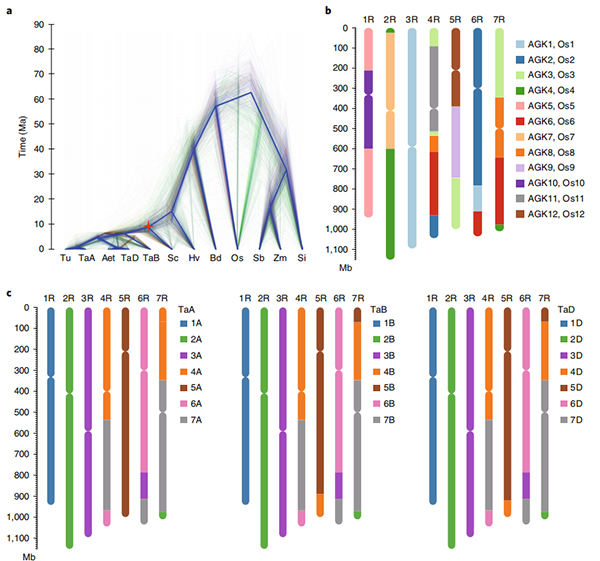 Nnyocha evolushọn na chromosome synteny nke rye genome |
Li, G., Wang, L., Yang, J.et al.Mgbakọ genome dị elu na-egosipụta njirimara genomic rye na mkpụrụ ndụ ihe nketa dị mkpa.Na Genet 53,574–584 (2021).
https://doi.org/10.1038/s41588-021-00808-z










