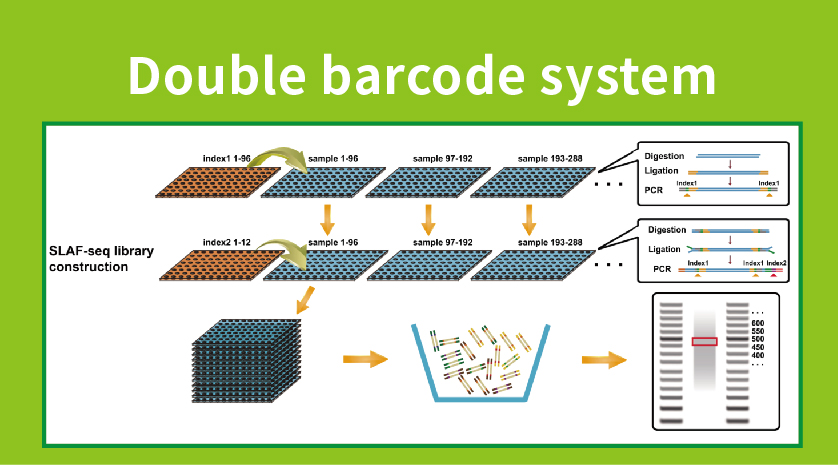
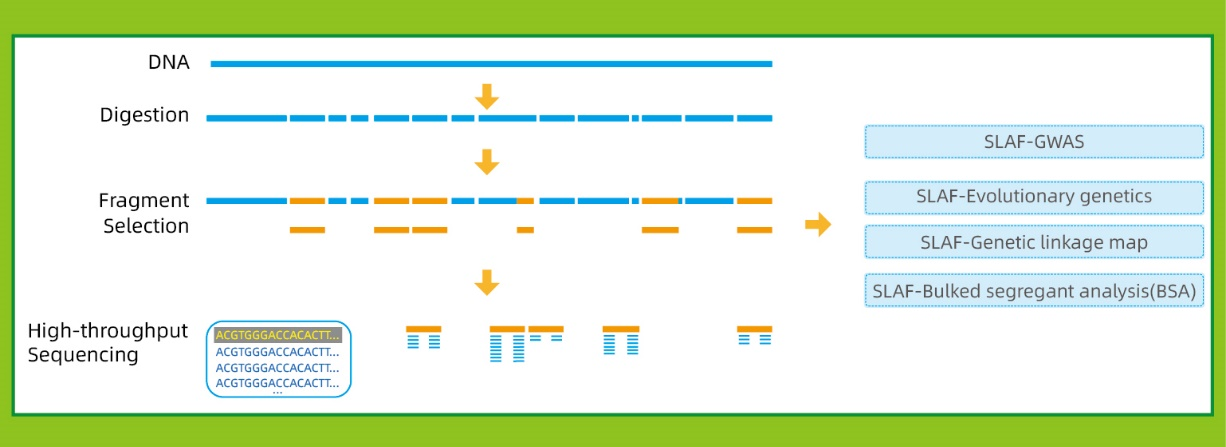
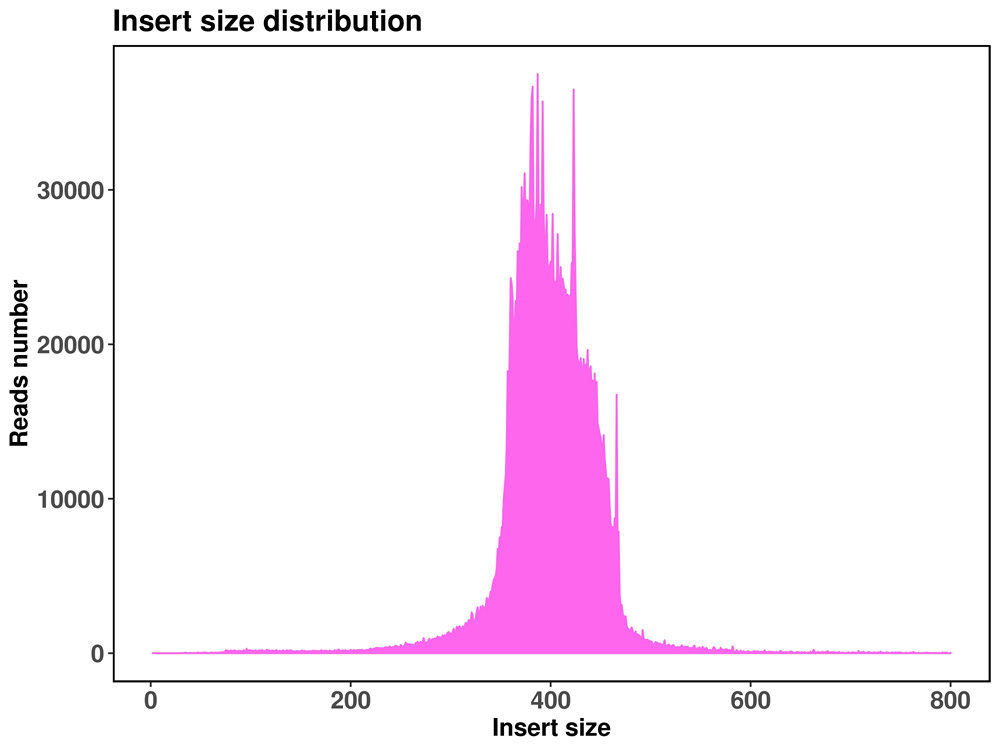
Usoro Mpempe Mpempe Akọwapụtara kpọmkwem-Locus (SLAF-Seq)
Nkọwa ọrụ
Atụmatụ nka
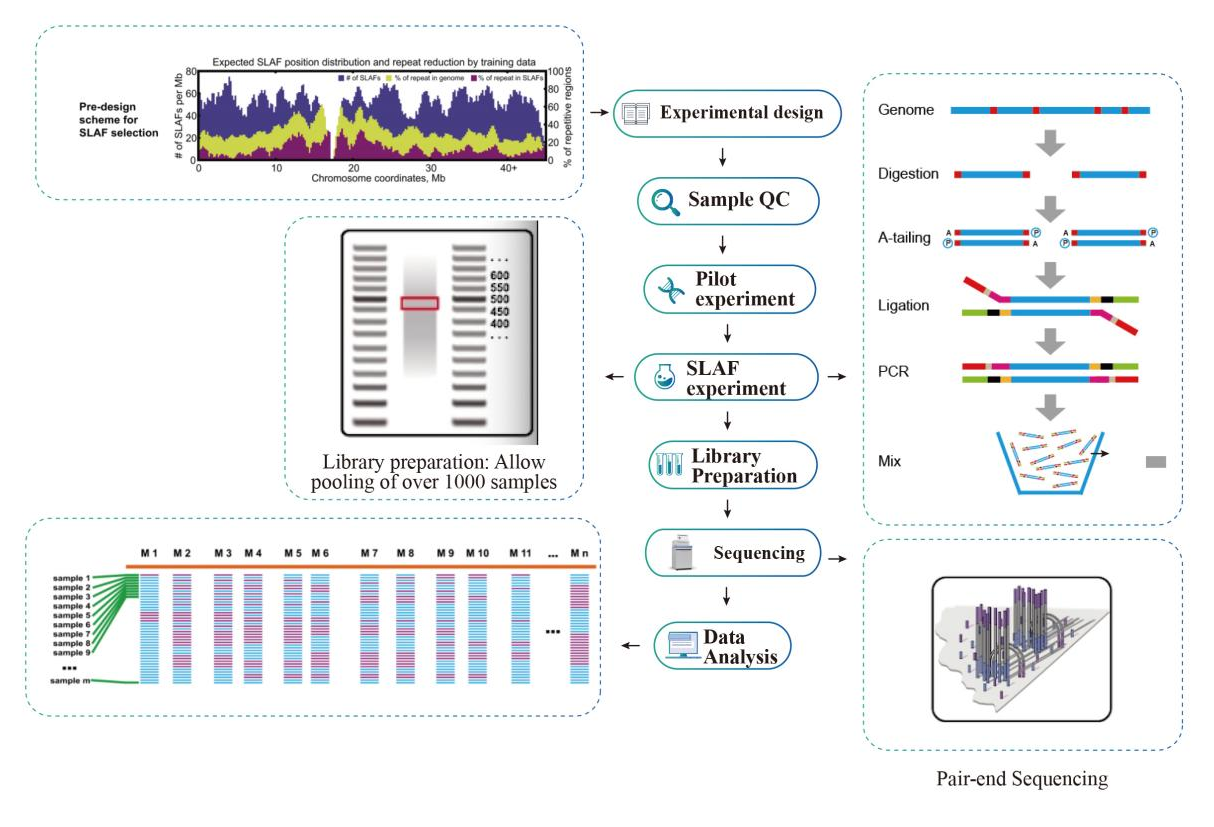
Usoro ọrụ

Uru ọrụ
Nchọpụta nrịbama dị elu- Teknụzụ usoro nrụpụta dị elu na-enyere SLAF-Seq aka ịchọpụta ọtụtụ narị puku mkpado n'ime genome niile.
Ndabere dị ala na genome- Enwere ike itinye ya na ụdị ma ọ bụ na-enweghị akwụkwọ genome.
Nhazi atụmatụ na-agbanwe agbanwe- Single-enzyme, dual-enzyme, multi-enzyme digestion na ụdị dị iche iche nke enzymes, niile nwere ike họrọ iji mee ihe dị iche iche nyocha ihe mgbaru ọsọ ma ọ bụ ụdị.A na-eji nyocha mbụ na silico iji mesie ike imepụta enzyme kacha mma.
Enzymatic mgbaze nke ọma- Emere nnwale tupu emee ka ọ dịkwuo mma, nke na-eme ka nnwale ahụ guzosie ike na ntụkwasị obi.Ịrụ ọrụ nchịkọta iberibe nwere ike nweta ihe karịrị 95%.
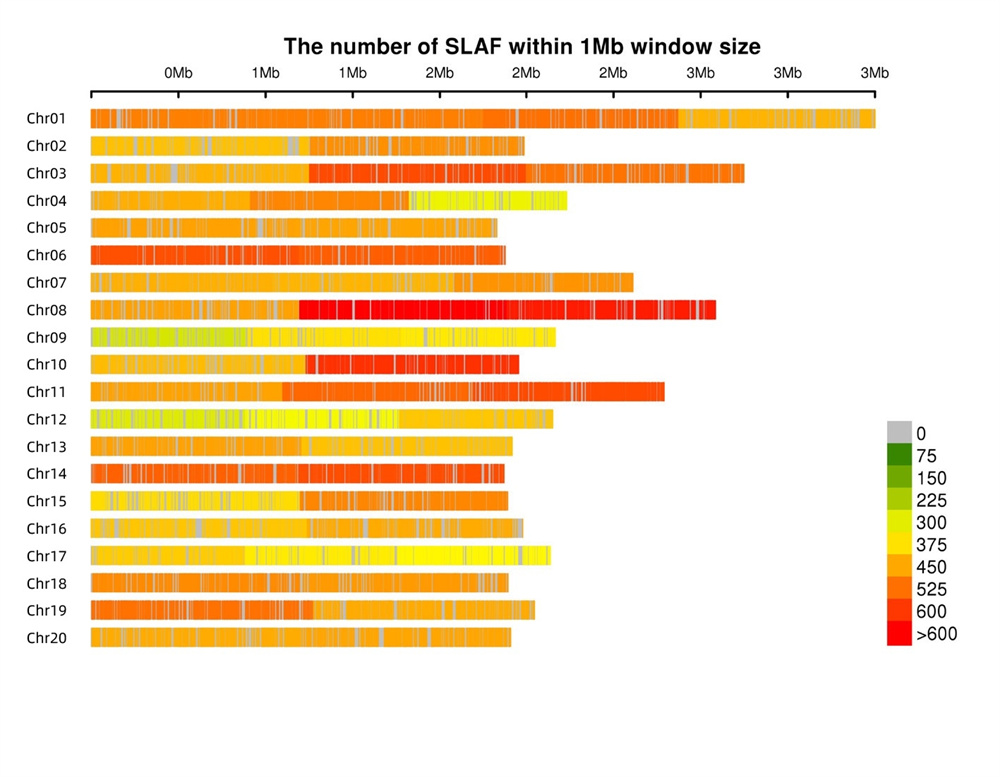
Mkpado SLAF ekesa nke ọma- A na-ekesa akara SLAF n'otu n'otu na chromosomes niile ruo oke, na-enweta nkezi 1 SLAF kwa 4 kb.
Mwepu nke ọma nke ikwugharị- A na-ebelata usoro ugboro ugboro na data SLAF-Seq ka ọ dị ala karịa 5%, ọkachasị n'ụdị nwere ọkwa dị elu nke ikwugharị, dị ka ọka wit, ọka, wdg.
Ahụmahụ sara mbara-N'ime 2000 mechiri emechi SLAF-Seq ọrụ na narị ụdị na-ekpuchi osisi, mammals, nnụnụ, ụmụ ahụhụ, aqua-ntule, wdg.
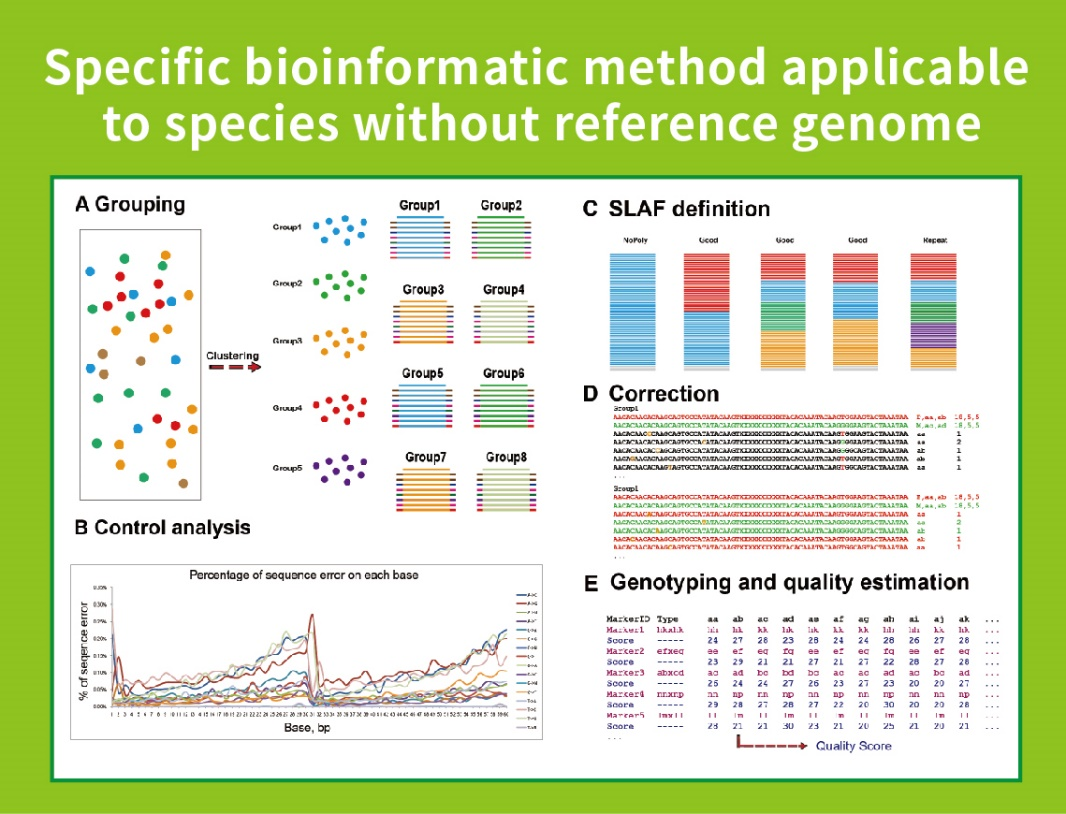
Nrụ ọrụ bioinformatic mebere onwe yaBMKGENE mepụtara usoro bioinformatic agbakwunyere maka SLAF-Seq iji hụ na ntụkwasị obi na izi ezi nke mmepụta ikpeazụ.
Nkọwapụta ọrụ
| Platform | Conc.(ng/gl) | Ngụkọta (ug) | OD260/280 |
| Illumina NovaSeq | >35 | >1.6(Mpịakọta>15μl) | 1.6-2.5 |
Atụmatụ usoro akwadoro
omimi usoro: 10X/Tag
| Nha genome | Mkpado SLAF akwadoro |
| <500 Mb | 100K ma ọ bụ WGS |
| 500 Mb-1 Gb | 100 K |
| 1 Gb -2 Gb | 200 K |
| Nnukwu ma ọ bụ mgbagwoju genome | 300-400K |
| Ngwa
| Akwadoro Ọnụ ọgụgụ ndị mmadụ
| Usoro nhazi usoro na omimi
| |
| Omimi
| Nọmba mkpado
| ||
| GWAS
| Nọmba nlele ≥200
| 10X
|
Dabere na genome size
|
| Genetic Evolution
| Ndị mmadụ n'otu n'otu otu obere ≥ 10; ngụkọta sample ≥ 30
| 10X
| |
Nnyefe nlele akwadoro
Akpa: 2 ml centrifuge tube
Maka ọtụtụ ihe atụ, anyị na-akwado ka ị ghara ichekwa na ethanol.
Ịkpọ aha nlele: Ọ dị mkpa ka e depụta ihe nlere anya nke ọma yana otu n'ụdị ozi nlele etinyere.
Mbupu: ice-akọrọ: A ga-ebu ụzọ tinye ihe nlele n'ime akpa wee lie ya na ice-akọrọ.
Usoro ọrụ ọrụ





Ihe atụ QC
Nnwale pilot
SLAF-nnwale
Nkwadebe ụlọ akwụkwọ
Usoro
Nyocha data
Ọrụ ire ere
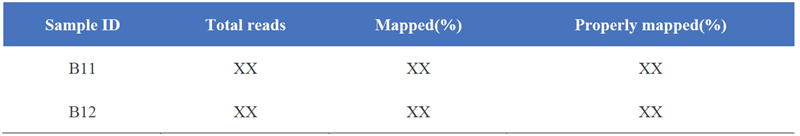
1. Statistics nke nsonaazụ map
2. mmepe akara SLAF
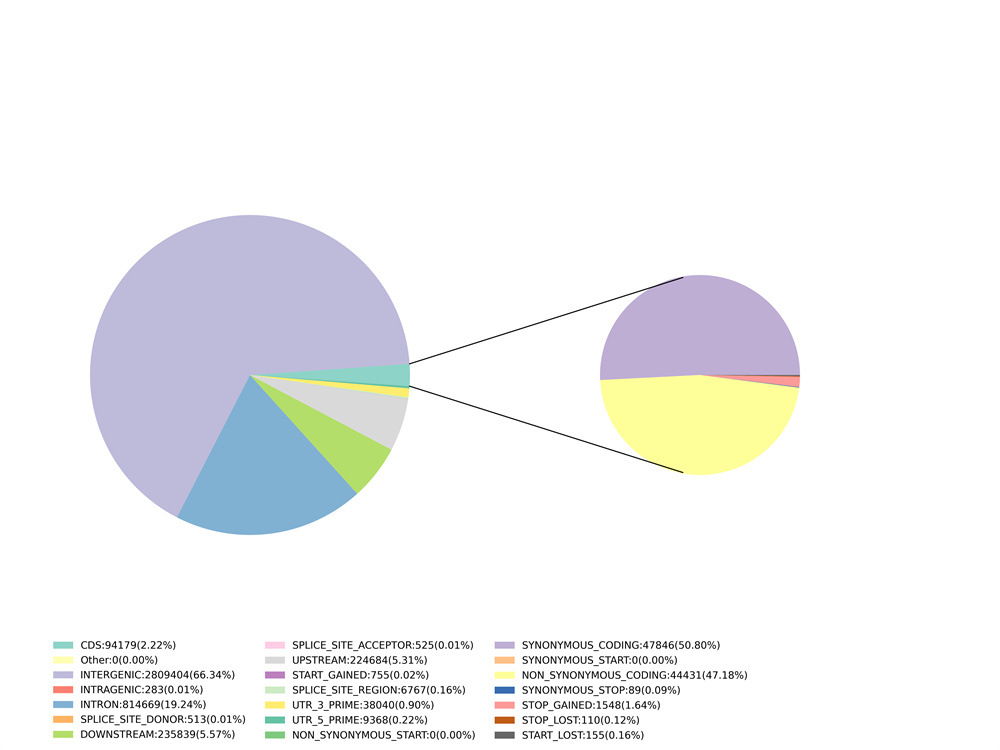
3. Nkọwa dị iche iche
| Afọ | Akwụkwọ akụkọ | IF | Aha | Ngwa |
| 2022 | Nkwukọrịta okike | 17.694 | Genomic ndabere nke giga-chromosomes na giga-genome nke osisi peony Paeonia ostii | SLF-GWAS |
| 2015 | Ọkachamara gbasara ahụike ọhụrụ | 7,433 | Akara ukwu nke ụlọ na-akwado mpaghara genomic nke mkpa agronomic dị na soybean | SLF-GWAS |
| 2022 | Akwụkwọ akụkọ nyocha dị elu | 12.822 | Genom-wide artificial introgressions nke Gossypium barbadense n'ime G. hirsutum kpughee loci dị elu maka imeziwanye ogo na mkpụrụ nke eriri owu n'otu oge àgwà | SAF-Evolutionary genetics |
| 2019 | Osisi Molecular | 10.81 | Ọnụọgụ Genomic Analysis na De Novo Assembly na-ekpughe mmalite nke ahịhịa Rice dị ka egwuregwu evolushọn | SAF-Evolutionary genetics |
| 2019 | Genetics okike | 31.616 | Usoro genome na ụdị mkpụrụ ndụ ihe nketa dị iche iche nke carp nkịtị, Cyprinus carpio | SLF-njikọ map |
| 2014 | Genetics okike | 25,455 | Mkpụrụ ndụ ihe nketa nke ahụekere na-akọ na-enye nghọta na legume karyotypes, polyploid evolushọn na ihe ubi domestication. | SLF-njikọ map |
| 2022 | Akwụkwọ akụkọ ihe ọkụkụ nke biotechnology | 9,803 | Nchọpụta nke ST1 na-ekpughe nhọrọ metụtara ịkụ mkpụrụ morphology mkpụrụ na mmanụ dị n'ime ya n'oge anụ ụlọ soybean | SLF-Marker mmepe |
| 2022 | Akwụkwọ akụkọ International nke Science Molecular | 6.208 | Nchọpụta na Mmepe Ihe nrịbama DNA maka Wheat-Leymus mollis 2Ns (2D) Ngbanwe chromosome Disomik | SLF-Marker mmepe |