PacBio 2+3 Full-Length mRNA Solution
Features
● Study design:
Pooled sample sequenced with PacBio to identify transcript isoforms
Separate samples (replicates and conditions to be tested) sequenced with NGS to quantify transcript expression
● PacBio sequencing in CCS mode, generating HiFi reads
● Sequencing of the full-length transcripts
● Analysis does not necessitate a reference genome; however, it may be employed
● Bioinformatic analysis includes not only expression at gene and isoform-level but also analysis of lncRNA, gene fusions, poly-adenylation, and gene structure
Advantages
● High accuracy: HiFi reads with accuracy >99.9% (Q30), comparable to NGS
● Alternative splicing analysis: sequencing all transcripts enables isoform identification and characterization.
● Combination of PacBio and NGS strengths: enabling quantification of expression at the isoform level, unveiling change that may be masked when analyzing the whole gene expression
● Extensive Expertise: with a track record of completing over 450 PacBio full-length transcriptome projects and processing over 700 samples, our team brings a wealth of experience to every project.
● Post-Sales Support: our commitment extends beyond project completion with a 3-month after-sale service period. During this time, we offer project follow-up, troubleshooting assistance, and Q&A sessions to address any queries related to the results.
Sample Requirements and Delivery
|
Library |
Sequencing strategy |
Data recommended |
Quality Control |
|
PolyA enriched mRNA CCS library |
PacBio Sequel II |
20 Gb |
Q30≥85% |
|
Poly A enriched |
Illumina PE150 |
6-10 Gb |
Q30≥85% |
Nucleotides
|
|
Conc.(ng/μl) |
Amount (μg) |
Purity |
Integrity |
|
Illumina Library |
≥ 20 |
≥ 0.5 |
OD260/280=1.7-2.5 OD260/230=0.5-2.5 Limited or no protein or DNA contamination shown on gel. |
For plants: RIN≥6.0; For animals: RIN≥6.5; 5.0≥28S/18S≥1.0; limited or no baseline elevation |
|
PacBio library |
≥ 100 |
≥ 0.75 |
OD260/280=1.7-2.5 OD260/230=0.5-2.5 Limited or no protein or DNA contamination shown on gel. |
Plants: RIN≥7.5 Animals: RIN≥8.0 5.0≥28S/18S≥1.0; limited or no baseline elevation |
Recommended Sample Delivery
Container: 2 ml centrifuge tube (Tin foil is not recommended)
Sample labeling: Group+replicate e.g. A1, A2, A3; B1, B2, B3.
Shipment
Dry-ice: Samples need to be packed in bags and buried in dry-ice.
RNAstable tubes: RNA samples can be dried in RNA stabilization tube(e.g. RNAstable®) and shipped in room temperature.
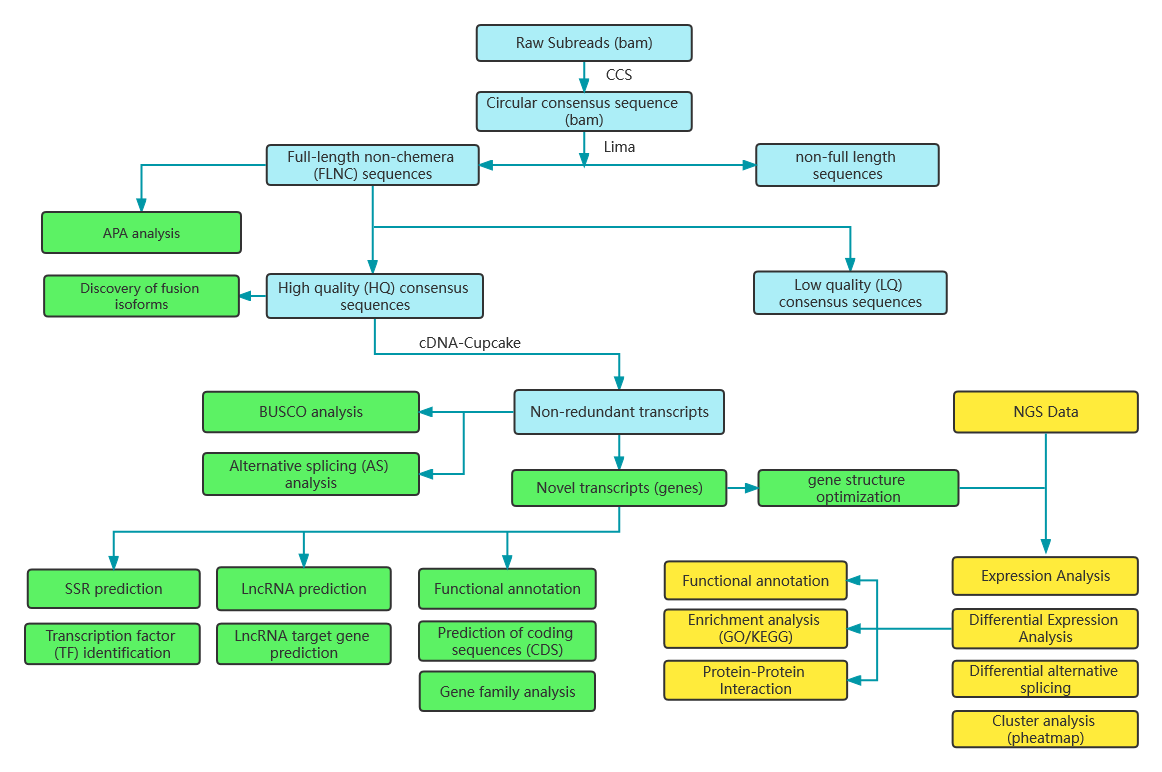
Includes the following analysis:
Raw data quality control
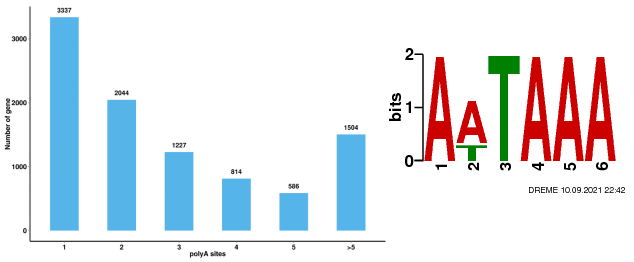
Alternative Polyadenylation Analysis (APA)
Fusion transcript analysis
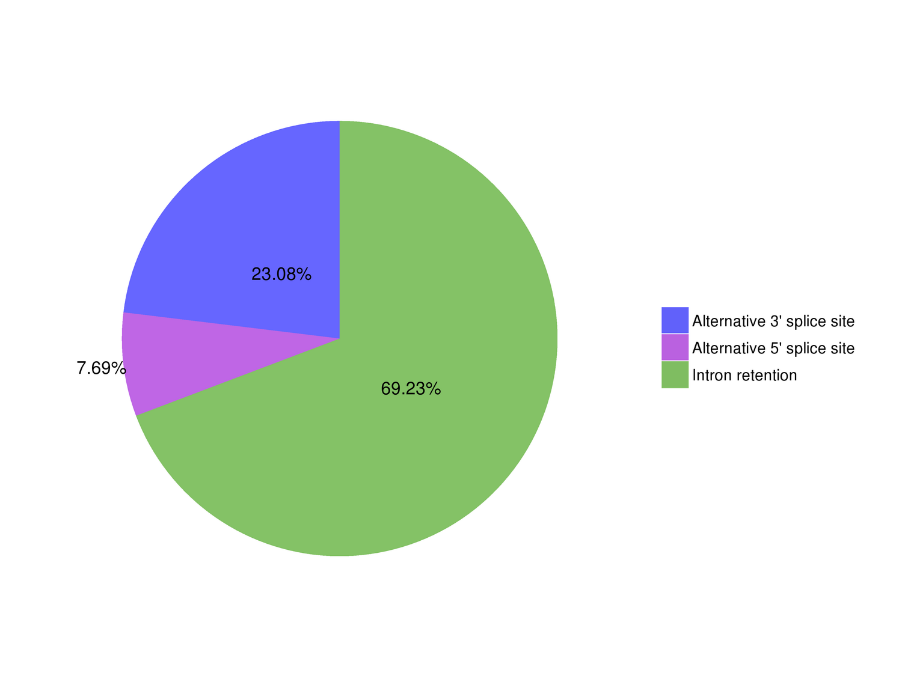
Alternative Splicing Analysis
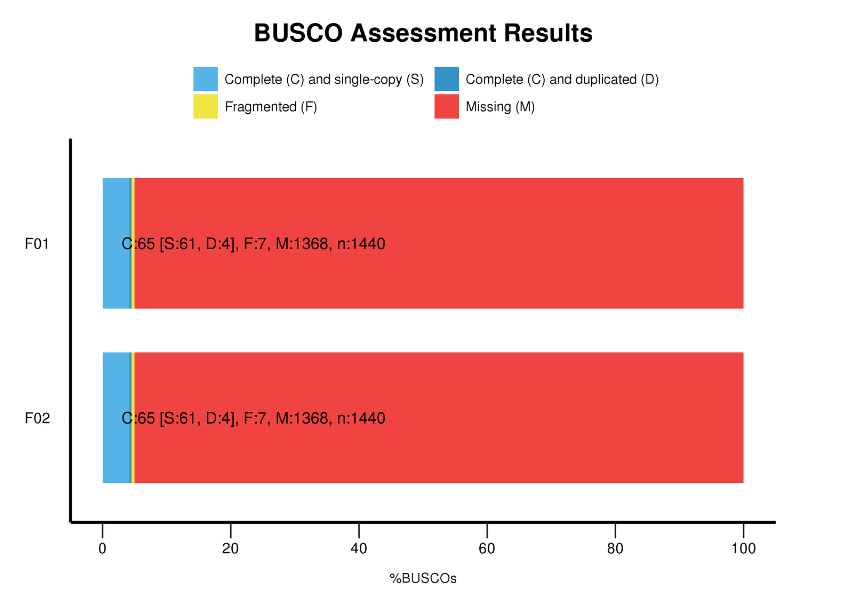
Benchmarking Universal Single-Copy Orthologs (BUSCO) analysis
Novel transcript analysis: prediction of coding sequences (CDS) and functional annotation
lncRNA analysis: prediction of lncRNA and targets
MicroSatelite Identification (SSR)
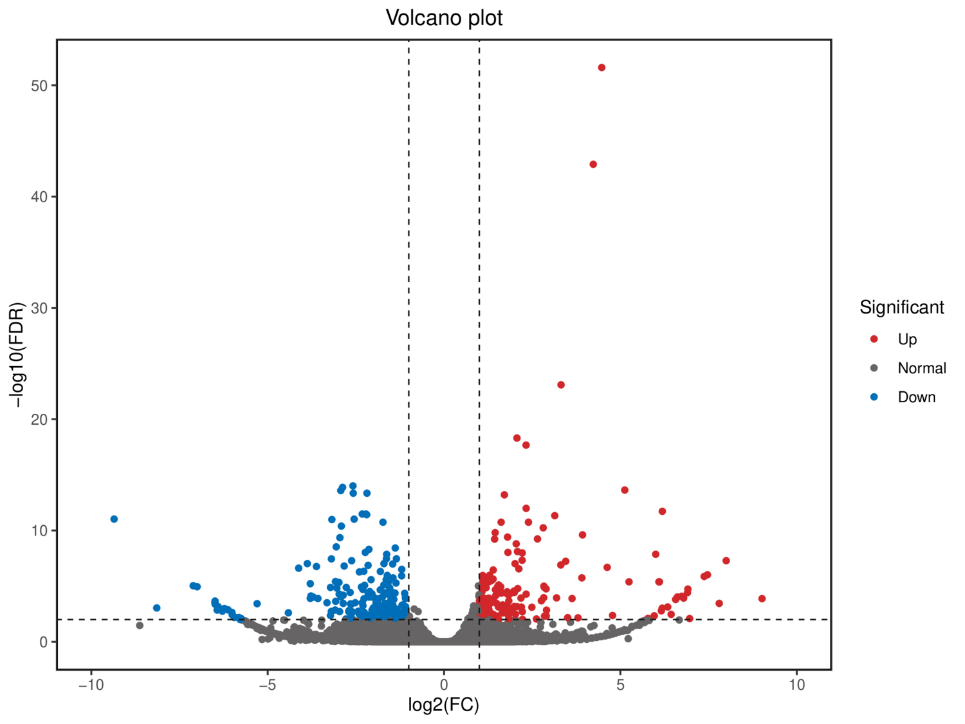
Differentially Expressed Transcripts (DETs) analysis
Differentially Expressed Genes (DEGs) analysis
Functional annotation of DEGs and DETs
BUSCO analysis
Alternative Splicing Analysis
Alternative Polyadenylation Analysis (APA)
Differentially Expressed Genes (DEGs) and Transcripts (DETs9 anlaysis
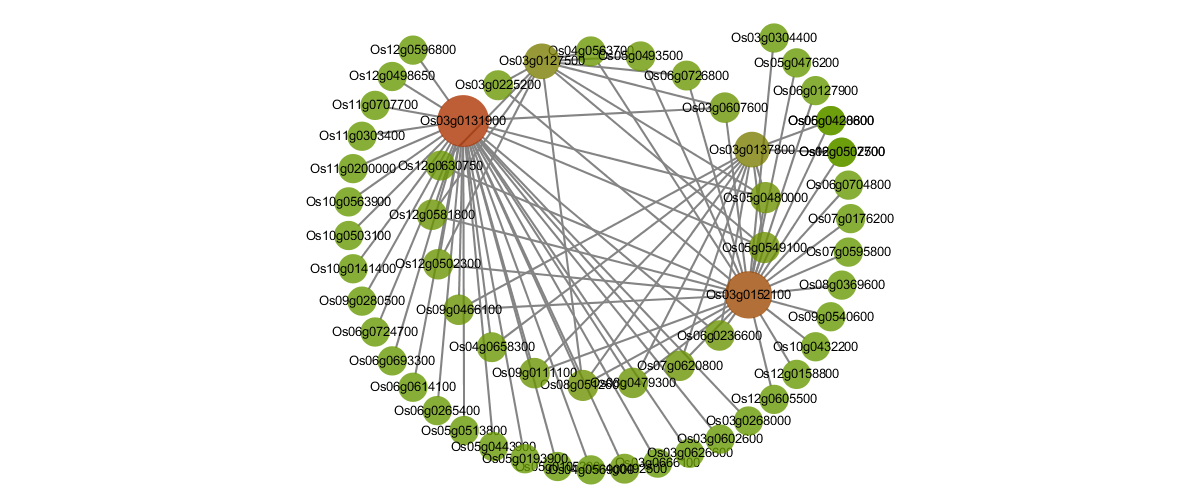
Protein-Protein interaction networks of DETs and DEGs
Explore the advancements facilitated by BMKGene’s PacBio 2+3 full-length mRNA sequencing through a curated collection of publications.
Chao, Q. et al. (2019) ‘The developmental dynamics of the Populus stem transcriptome’, Plant Biotechnology Journal, 17(1), pp. 206–219. doi: 10.1111/PBI.12958.
Deng, H. et al. (2022) ‘Dynamic Changes in Ascorbic Acid Content during Fruit Development and Ripening of Actinidia latifolia (an Ascorbate-Rich Fruit Crop) and the Associated Molecular Mechanisms’, International Journal of Molecular Sciences, 23(10), p. 5808. doi: 10.3390/IJMS23105808/S1.
Hua, X. et al. (2022) ‘Effective prediction of biosynthetic pathway genes involved in bioactive polyphyllins in Paris polyphylla’, Communications Biology 2022 5:1, 5(1), pp. 1–10. doi: 10.1038/s42003-022-03000-z.
Liu, M. et al. (2023) ‘Combined PacBio Iso-Seq and Illumina RNA-Seq Analysis of the Tuta absoluta (Meyrick) Transcriptome and Cytochrome P450 Genes’, Insects, 14(4), p. 363. doi: 10.3390/INSECTS14040363/S1.
Wang, Lijun et al. (2019) ‘A survey of transcriptome complexity using PacBio single-molecule real-time analysis combined with Illumina RNA sequencing for a better understanding of ricinoleic acid biosynthesis in Ricinus communis’, BMC Genomics, 20(1), pp. 1–17. doi: 10.1186/S12864-019-5832-9/FIGURES/7.