-

Metagenomic Sequencing -NGS
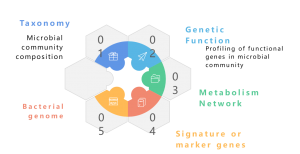
Metagenome refers to a collection of the total genetic material of a mixed community of organisms, such as environmental and human metagenomes. It contains genomes of both cultivatable and uncultivatable microorganisms. Shotgun metagenomic sequencing with NGS enables the study of these intricate genomic landscapes embedded in environmental samples by providing more than taxonomic profiling, giving also granular insights into species diversity, abundance dynamics, and complex population structures. Beyond taxonomic studies, shotgun metagenomics also offers a functional genomics perspective, enabling the exploration of encoded genes and their putative roles in ecological processes. Finally, the establishment of correlation networks between genetic elements and environmental factors contributes to a holistic understanding of the intricate interplay between microbial communities and their ecological background. In conclusion, metagenomic sequencing stands as a pivotal instrument for unravelling the genomic intricacies of diverse microbial communities, illuminating the multifaceted relationships between genetics and ecology within these complex ecosystems.
Platforms: Illumina NovaSeq and DNBSeq T7
-

Metagenomic Sequencing-Nanopore
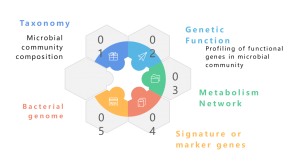
Metagenome refers to a collection of the total genetic material of a mixed community of organisms, such as environmental and human metagenomes. It contains genomes of both cultivatable and uncultivatable microorganisms. Metagenomic sequencing enables the study of these intricate genomic landscapes embedded in environmental samples by providing more than taxonomic profiling, also offering a functional genomics perspective by exploring the encoded genes and their putative roles in ecological processes. While traditional shotgun approaches with Illumina sequencing, have been widely used in metagenomic studies, the advent of Nanopore long-read sequencing has changed the field. Nanopore technology enhances downstream bioinformatic analyses, notably metagenome assembly, ensuring more continuous assemblies. Remarkably, reports indicate that Nanopore-based metagenomics has successfully generated complete and closed bacterial genomes from complex microbiomes (Moss, E. L., et. al, Nature Biotech, 2020). Integrating Nanopore reads with Illumina reads provides a strategic approach for error correction, mitigating Nanopore’s inherent low accuracy. This synergistic combination leverages the strengths of each sequencing platform, offering a robust solution to overcome potential limitations and advancing the precision and reliability of metagenomic analyses.
-

16S/18S/ITS Amplicon Sequencing-PacBio
The 16S and 18S rRNA genes, along with the Internal Transcribed Spacer (ITS) region, serve as pivotal molecular fingerprinting markers due to their combination of highly conserved and hyper-variable regions, making them invaluable tools for characterizing prokaryotic and eukaryotic organisms. Amplification and sequencing of these regions offer an isolation-free approach for investigating the microbial composition and diversity across various ecosystems. While Illumina sequencing typically targets short hypervariable regions like V3-V4 of 16S and ITS1, it has been demonstrated that superior taxonomic annotation is achievable by sequencing the full length of 16S, 18S, and ITS. This comprehensive approach results in higher percentages of accurately classified sequences, achieving a level of resolution that extends to species identification. PacBio’s Single-Molecule Real-Time (SMRT) sequencing platform stands out by providing highly accurate long reads (HiFi) that cover the full-length amplicons, rivalling the precision of Illumina sequencing. This capability allows researchers to attain an unmatched advantage — a panoramic view of the genetic landscape. The extended coverage significantly elevates the resolution in species annotation, particularly within bacterial or fungal communities, enabling a deeper understanding of the intricacies of microbial populations.
-

16S/18S/ITS Amplicon Sequencing-NGS
Amplicon sequencing with Illumina technology, specifically targeting the 16S, 18S, and ITS genetic markers, is a powerful method for unraveling the phylogeny, taxonomy, and species abundance within microbial communities. This approach involves sequencing the hypervariable regions of housekeeping genetic markers. Originally introduced as a molecular fingerprint by Woeses et al in 1977, this technique has revolutionized microbiome profiling by enabling isolation-free analyses. Through the sequencing of 16S (bacteria), 18S (fungi), and Internal Transcribed Spacer (ITS, fungi), researchers can identify not only abundant species but also rare and unidentified ones. Widely adopted as a pivotal tool, amplicon sequencing has become instrumental in discerning differential microbial compositions across diverse environments, including the human mouth, intestines, stool, and beyond.
-

Bacterial and Fungal Whole Genome Re-Sequencing
Bacterial and fungal whole-genome re-sequencing projects are pivotal for advancing microbial genomics, by enabling the completion and comparison of microbial genomes. This facilitates fermentation engineering, the optimization of industrial processes, and the exploration of secondary metabolism pathways. Furthermore, fungal and bacterial re-sequencing is crucial for understanding environmental adaptation, optimizing strains, and revealing genetic evolution dynamics, with broad implications in medicine, agriculture, and environmental science.
-

Metatranscriptome Sequencing
Leveraging Illumina sequencing technology, BMKGENE’s metatranscriptome sequencing service unveils the dynamic gene expression of a diverse array of microbes, spanning eukaryotes to prokaryotes and viruses, within natural environments such as soil, water, sea, stool, and the gut. Our comprehensive service empowers researchers to delve into the entirety of gene expression profiles within complex microbial communities. Beyond taxonomic analysis, our metatranscriptome sequencing service facilitates exploration into functional enrichment, shedding light on differentially expressed genes and their roles. Uncover a wealth of biological insights as you navigate the complex landscapes of gene expression, taxonomic diversity, and functional dynamics within these diverse environmental niches.
-

Fungal complete genome
BMKGene offers versatile solutions for fungal genomes, catering to diverse research needs and desired genome completeness. Utilizing short-read Illumina sequencing alone allows the generation of a draft genome. For a more refined fungal genome with longer contigs, both short-reads and long-read sequencing using Nanopore or PacBio are combined. Moreover, the integration of Hi-C sequencing further enhances the capabilities, enabling the attainment of a complete chromosome-level genome.
-

Bacterial complete genome
BMKGene provides a complete bacterial genome assembly service, guaranteeing 0 gaps. This is made possible by the integration of long-read sequencing technologies Nanopore and PacBio for assembly and short-read sequencing with Illumina for assembly validation and error correction of ONT reads. Our service provides the complete bioinformatic workflow from assembly, functional annotation, and advanced bioinformatic analysis, fulfilling specific research goals. This service enables the development of precise reference genomes for various genetic and genomic studies. Additionally, it forms the basis for applications such as strain optimization, genetic engineering, and microbial technology development, ensuring reliable and gap-free genomic data crucial for advancing scientific insights and biotechnological innovation.